MicroRNAs (miRNAs) are non-coding RNAs containing 18-25 nucleotides encoded by endogenous genes and have the ability to regulate gene expression at the post-transcriptional level by repressing mRNA or promoting mRNA degradation. Typically, genes encoding miRNAs are more conserved in species that are very closely related, but they are sometimes homologous in species that are very distantly related.
Various physiological processes (hematopoiesis, organogenesis, lipid metabolism, etc.) and pathologies (including cancer, cardiovascular and metabolic diseases) are highly correlated with the expression patterns of miRNA. However, their activity does not always correlate with the expression levels. Several mechanisms have been reported to influence the action and function of miRNA, which include genetic polymorphisms, miRNA strand selection, interactions with RNA binding proteins (RBPs) or other coding/non-coding RNAs.
There are three main effects of miRNAs in animals, (1) inhibiting translation of target genes, (2) inducing mRNA cleavage or (3) degradation.
The inhibition translation of mRNA by miRNAs can occur at two stages: before transcription initiation, or at the stage of translation extension.
The cleavage of mRNA is not common in animals. There are two conditions that need to be met before the mRNA is cleaved, the miRNA matches the target gene as perfectly as possible, and the miRNA is wrapped by AGO2.
When the transcription of mRNA is inhibited, miRNA may trigger mRNA deadenylation and decapping, which then causes mRNA degradation, and this may be the main way to reduce mRNA degradation caused by miRNA in animals.
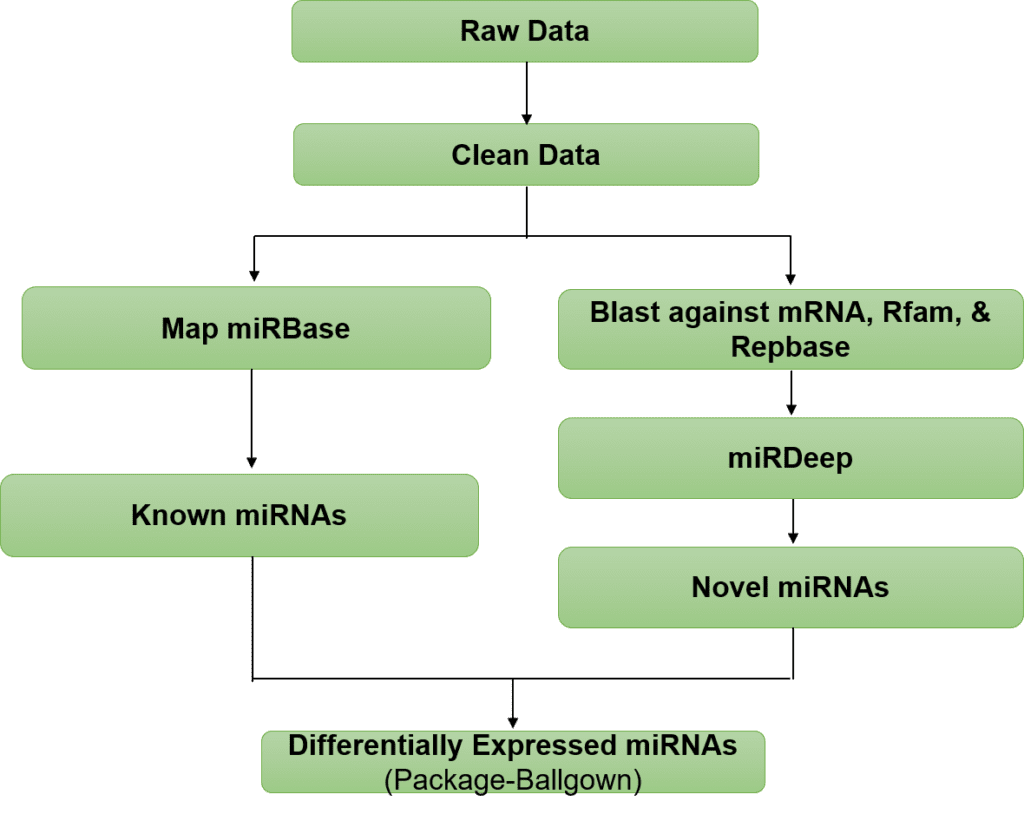
DNA/RNA sequencing, a new generation of high-throughput technologies capable of identifying molecules with great sensitivity, specificity and predictive power to detect disease, is also being applied to the detection and analysis of miRNAs. The use of miRNA sequencing has two main advantages. One is allowing the discovery of novel miRNAs, and the other is the utilization of molecular barcodes allowing for accurate miRNA quantification. Combining RNA immunoprecipitation and next generation sequencing, RIP-Seq analyzes the interaction between miRNAs and proteins.
In addition, nucleotide modifications are critical for miRNA. These modifications include m6A, m7G, and m5C, etc. The modified miRNAs modulate the stability and translation efficiency of target genes by regulating miRNA-mRNA or miRNA-protein interactions. miRNA m6A can be comprehensively detected by MeRIP-seq, and m5C can be comprehensively analyzed by hMeRIP-seq, m5C-RIP-seq and RNA BS-seq.
Recently, nanopore sequencing based methods for detecting miRNA expression patterns have been developed that are capable of identifying cancer-specific expression patterns from samples with very low miRNA concentrations. Nanopore decoding technology is expected to be a simple and biomarker discovery tool.
miRNAs are naturally occurring molecules in humans and can target multiple genes at the same time. The activity of miRNAs can be greatly influenced by altering the specificity of miRNAs to target mRNAs. Therefore, it is widely used in the fields of disease research and the development of microRNA therapeutics.
The miRNA expression profile of tumor tissues or organs is altered to present a disease-specific expression profile. Taking advantage of this, new biomarkers can be discovered by microRNA sequencing, which can help develop cancer screening and diagnostic technologies.
Studies have shown that many miRNAs are closely related to the development of several diseases, including cancer. Therefore, controlling the expression of these cancer-related miRNAs is a new cancer treatment strategy. Currently, there are two main types of miRNA therapies: 1) miRNA inhibitors, and 2) miRNA mimics (alternative therapies).